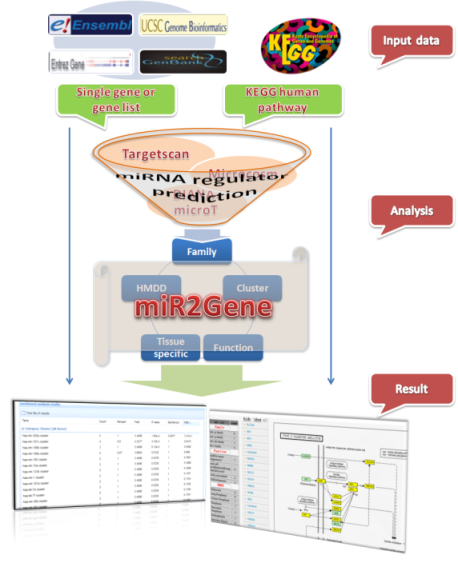
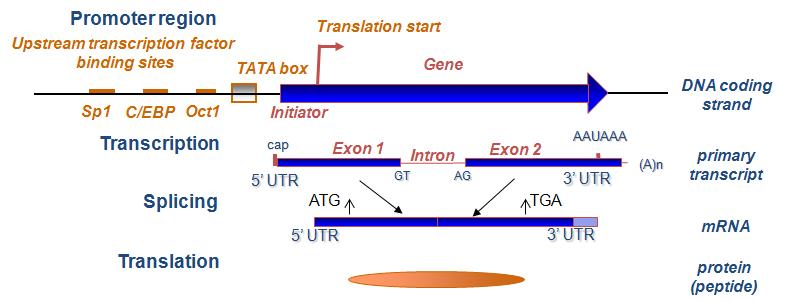
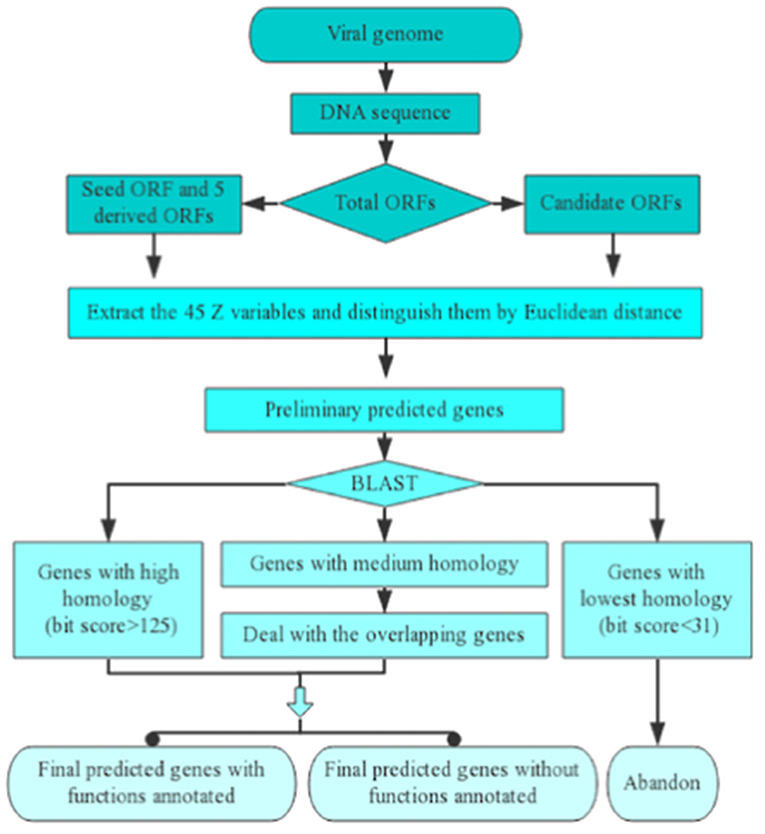
Schematic representation of the various tools of the genome annotation... | Download Scientific Diagram
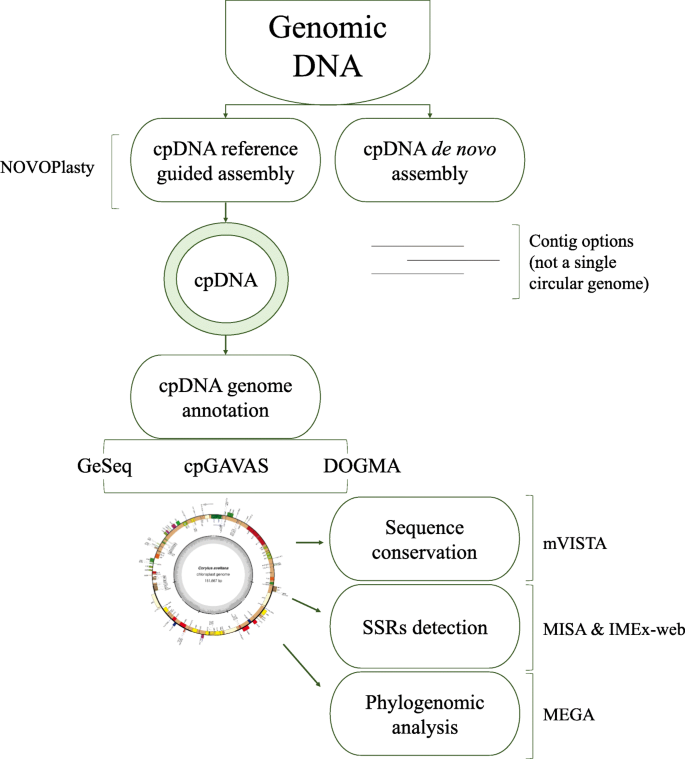
Comparison of different annotation tools for characterization of the complete chloroplast genome of Corylus avellana cv Tombul | BMC Genomics | Full Text
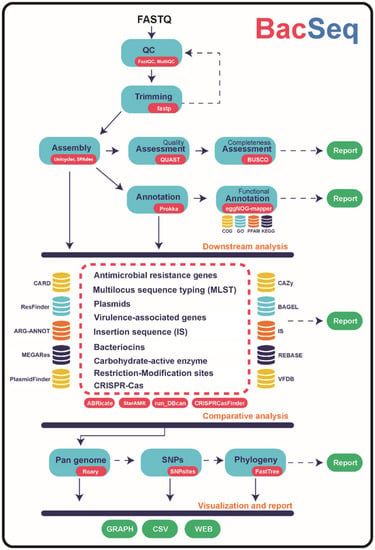
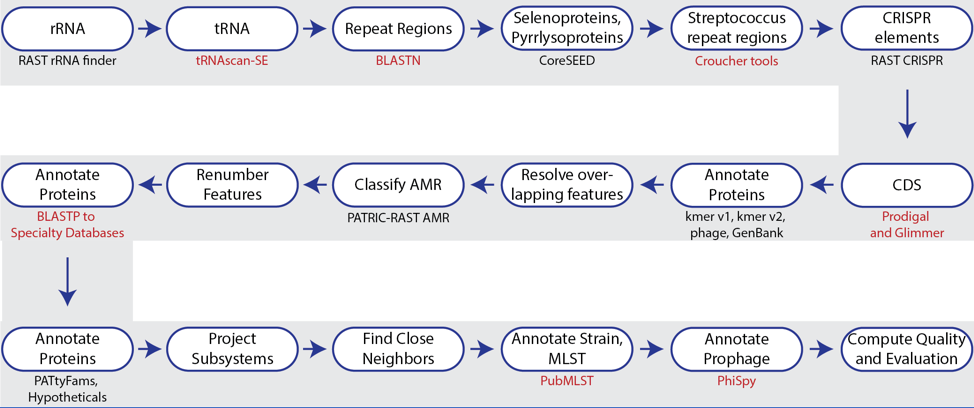
Microorganisms | Free Full-Text | BacSeq: A User-Friendly Automated Pipeline for Whole-Genome Sequence Analysis of Bacterial Genomes

KEMET – A python tool for KEGG Module evaluation and microbial genome annotation expansion - Computational and Structural Biotechnology Journal

GitHub - galikbence/genome_annotation: Pipeline for eukaryotic genome annotation based on external evidences

Genome Research on X: "@genomeresearch publishes a Perspective focusing on the expression of alternative ORFs, the current tools for studying them, and proposes a genome annotation framework that includes alternative ORFs to
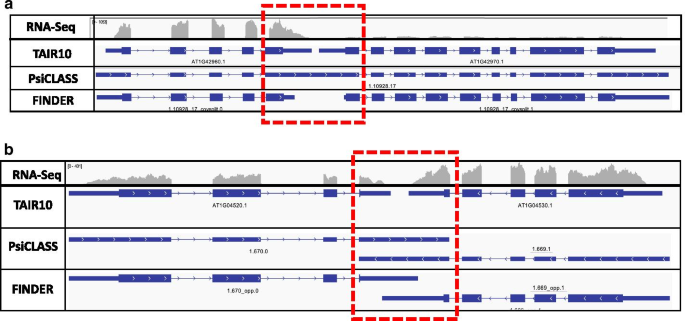
FINDER: an automated software package to annotate eukaryotic genes from RNA-Seq data and associated protein sequences | BMC Bioinformatics | Full Text
![PDF] Review on the Computational Genome Annotation of Sequences Obtained by Next-Generation Sequencing | Semantic Scholar PDF] Review on the Computational Genome Annotation of Sequences Obtained by Next-Generation Sequencing | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/4d901834206df869fcf2e8a144913ae799925a6d/7-Table2-1.png)
PDF] Review on the Computational Genome Annotation of Sequences Obtained by Next-Generation Sequencing | Semantic Scholar







![PDF] Annotation, comparison and databases for hundreds of bacterial genomes. | Semantic Scholar PDF] Annotation, comparison and databases for hundreds of bacterial genomes. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/d9bfe8b9385f7cfdc2c78df99d1c39040c9a3e7b/3-Table1-1.png)